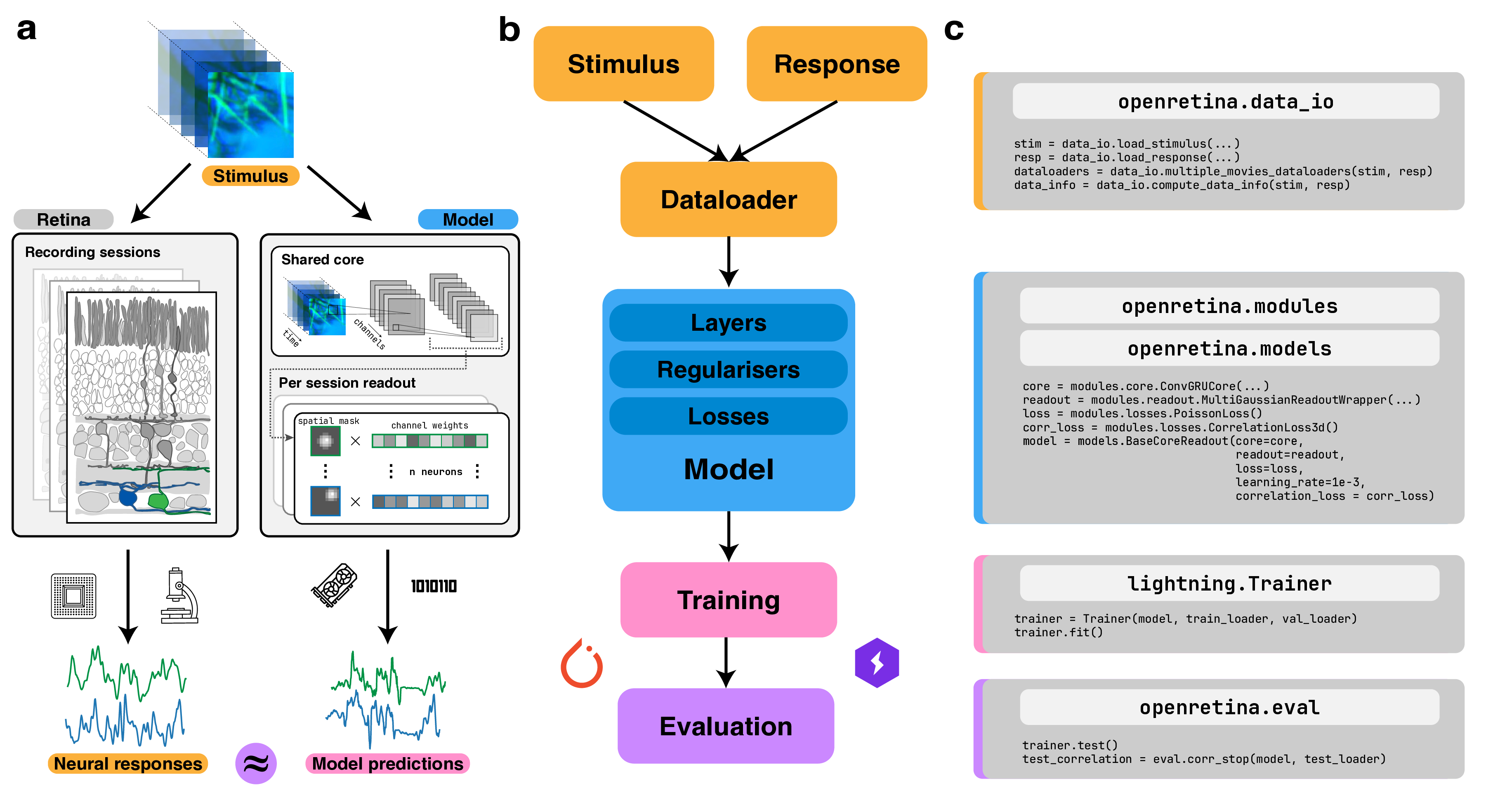
Training Overview
openretina builds on top of PyTorch, PyTorch Lightning, and Hydra to provide a flexible training stack. Lightning handles training loop infrastructure (logging, checkpoints, devices, precision), while Hydra keeps configuration modular and composable. This combination makes experiments reproducible and easier to share.
If you are preparing data for training, start with the Data I/O guide. When you are ready to run a full experiment, use the Unified Training Script. For a task-based map of notebook and CLI paths, see How to....
What the Training Infrastructure Provides
- Configuration management: reusable Hydra configs under
configs/, including presets for datasets, models, and optimisation hyper-parameters. - Experiment tracking: automatic logging of losses, metrics, and checkpoints via PyTorch Lightning, with optional integrations for TensorBoard and WandB.
- Modular model components: interchangeable cores, readouts, regularisers, and losses from
openretina.modules, orchestrated throughopenretina.models. - Evaluation utilities: trained model evaluation via
openretina.eval.
Typical Training Workflow
- Select data: Load stimuli and responses into dictionaries of train/validation/test splits as described in the Data I/O documentation.
- Choose configuration: Pick a base config (e.g.
configs/hoefling_2024_core_readout_low_res.yaml) and customise modules, hyper-parameters, or logging targets. - Launch training: Run
openretina train ...or the equivalent Python script. Lightning handles checkpoints, gradient accumulation, and mixed precision when enabled. - Monitor progress: Inspect metrics in TensorBoard or your preferred logger. Checkpoints are saved under the run directory specified by Hydra.
- Evaluate results: Use
openretina evaland/or notebooks to benchmark against ground truth and inspect model behavior.
The notebooks in notebooks/ mirror these steps with executable examples, including reproductions of the core + readout model on both calcium imaging and spiking datasets.